GIMP stand for GNU Image Manipulating Program. I have this software in my computer on and off for the last few years. I was adamant to not budge for paid softwares like Adobe photo shop and like. Gimp used to sit there in my computer and annoy me with its small icon, so I used to uninstall it often. When I needed to manipulate some figures for publication, I would install it back on. Not until very recently I had no clue how powerful and useful gimp can be. It can even beat the best image editing softwares in market..
Challenges:
Recently, I faced a challenge merging 3 pathway maps. They can be as complicated as a circuit diagram!! I had to basically merge Image 1 and Image 2 and make a composite image12. Then I had to merge this with image 3 in such a way that, imae3 - image12 will be in a different color and image12 - image3 will be in a different color.
Open image 1 as layer(file --> open as layer), then open image 2 as layer as well. Then go to layer menu --> Merge down --> save image as image12. In image12, you will see the extra portions of image 1 as little lighter objects, but image 2 appears as it is. Now open image12 as layer then open image 3 as layer. Then go to layer menu --> merge down --> save as image123.
Now in image123 you see the extra portions in 12 that is not present in 3 as lighter objects. Well enough..
Now do the reverse. Open image 3 as layer first followed by image 12 as layer. Then merge as before and name it as image 312.
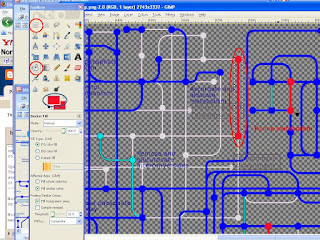
What you will see is in image 312 --> All that is present extra in 3 are not present in image 12 will appear as solid bordered transparent object. Distinct from the upper layer. In Image 123 you will see just the reverse. The ones that are present extra in image 12 and are not present in image 3. Now after all these, you can happily open image 312 and start playing around with colors. Open image 312 and as shown in figure select the shape from the toolbox menu. Select the area neatly that need to be colored differently. Then choose color. Then select color menu and just click on the selected area in the image. Viola.. that changes the color so well....
